Combined Bacterial Antigen Lipopolysaccharide and Lipoteichoic Acid Increase Cal 27 Oral Cancer Cell Proliferation
A B S T R A C T
Oral biofilms harbour gram-negative bacterial antigen lipopolysaccharide (LPS) involved in oral cancer progression and gram-positive bacterial surface-associated adhesive, lipoteichoic acid (LTA). Thus, we hypothesised that LPS and LTA together would increase the proliferation of cancer cells compared to stimulation by LPS alone. Oral cancer cell lines SCC4, SCC9, SCC25, Cal 27 and the normal oral cell line, OKF6, were studied. The bacterial antigen stimulation indices were determined using the MT Glo assay. Cell proliferation after bacterial antigen stimulation was validated by clonogenic assays. Phosphokinase array, reverse transcription-quantitative polymerase chain reaction (RT-qPCR), and Western blot were employed to study proliferative and apoptotic pathways in bacterial antigen-stimulated cells. Bacterial antigens significantly stimulated Cal 27 (p ≤ 0.001) alone. SCC4 and SCC9 showed negligible stimulation with either antigen, while SCC25 results were comparable to OKF6. The combined antigen stimulation of Cal 27 led to a decrease in phosphorylated p53 and β-catenin and higher PI3K compared to LPS only stimulated cells (p ≤ 0.001). Combined bacterial antigen stimulation results in increased proliferation of Cal 27 cells due to lowering of tumor suppressor proteins and increased tumor proliferation-related proteins.
Keywords
Bacterial antigen, LPS, LTA, biofilm, oral cancer cell, proliferation, phosphokinase
Introduction
Oral cancer is the sixth most common cancer in the world, high risk in South Asia with male preponderance [1, 2]. Lipopolysaccharides (LPS) are found in the cell wall of Gram-negative bacteria such as Porphyromonas gingivalis and Fusobacterium nucleatum, which take part in biofilm formation [3]. LPS has been reported to be associated with the progression of oral cancer [3]. Fusobacterium nucleatum LPS has been shown to upregulate β-catenin, which activates the c-myc, NF-kB and cyclin D kinases, resulting in the activation of cyclin D and consequent cancer cell survival and proliferation [4]. While P. gingivalis LPS downregulates p53 and upregulates NF-kB, MMP-9, and the JAK/STAT pathway, particularly STAT3 (which controls the intrinsic mitochondrial apoptotic pathway) [4]. This results in inflammation, anti-apoptosis, increased viability of oral cancer cells and increased oral cancer invasion [4]. Human hepatocyte growth factor (HuHGF) stimulates stromal fibroblast-induced invasion (epithelial-mesenchymal cells) in the progression of oral cancer [5]. HuHGF is upregulated by LTA from bacteria (Streptococcus mutans, Streptococcus pyogenes, Streptococcus faecalis, Streptococcus sanguis, and Streptococcus aureus) of oral biofilm [6]. Other than its induction of HuHGF, the role of LTA in the proliferation of oral cancer cells remains elusive [6]. Therefore, we hypothesised that LPS and LTA together would increase the proliferation of cancer cells compared to stimulation with LPS alone. We present here the effect of combined bacterial antigen on select oral cancer cell lines and explore the proliferation pathways that may be affected by these bacterial antigens.
Materials and Methods
Details of all materials and reagents used in this study are provided in (Supplementary Table 1).
I Culture and Maintenance of Cells
We used American Type Culture Collection (ATCC) cell lines SCC4 (CRL-1623), SCC9 (CRL-1629), SCC25 (CRL-1628), Cal 27 (CRL-2095), and OKF6 (CVCL_L222). Details are available in (Supplementary Table 2). Molecular profiling of the cell lines has shown that they can be used as preclinical models for oral cancer translational research [7]. Following ATCC guidelines, Cal 27, SCC4, and SCC9 were cultured using Dulbecco’s Modified Eagle Medium (DMEM) (Gibco, Cat #11960-044) plus 10% foetal bovine serum (FBS, Life Technologies, Cat #10099141) plus 1% penicillin (P)/streptomycin (S) (Life Technologies, Cat #15140-148). To culture OKF6 and SCC25, we used keratinocyte serum-free medium (KSFM, Life Technologies Cat #1700504) plus growth factors (Gibco, human recombinant epithelial growth factor (EGF), Cat #10450-013 bovine pituitary extract (BPE, Cat #13028-014). The medium for all the cell lines was changed twice every week. Each 75cm2 flask of cells acted as a biological replicate. Generally, the biological replicate was split into three technical replicates. Biological replicates i.e., flasks, were throughout from three different cell passages and conducted on different days (or at different time points). Throughout this report, n refers to the number of biological replicates.
II Stimulation Index
The stimulation index for each bacterial antigen was done following the Real Time-Glo MT Cell Viability Assay (Promega, Cat #G9712). 5000 cells were plated (96-well plate) for Cal 27, SCC25, and OKF6 and treated with 5 µg/ml LPS (from Escherichia coli O111:B4, Sigma Aldrich), LTA (from Streptococcus pyogenes, Sigma Aldrich), or the combination [LPS (5 µg/ml) +LTA (5 µg/ml)]. The luminescence of the treated cells was measured using a plate-reading luminometer (BMG Labtech microplate reader PHERAstar).
III Clonogenic Assay
Cal 27 at 5000 cells were plated into each well of a 24-well plate. The bacterial antigen treatments were performed for 72 hr on the cells. To fix the cells, 1 ml 100% ice-cold methanol (Chem-Supply, Cat #MA004-2.5L-P), was added. After 1 min, the methanol was removed and the plates were air-dried until completely dry, followed by the addition of 0.1% crystal violet (Sigma Aldrich, Cat #C5042), allowed to develop for 20 min.
IV Phosphokinase Array
Proteome Profiler Human Phospho-Kinase Array kit (R&D Technologies, Cat #ARY003C) was used to identify proliferation-related phosphorylated protein expression in the bacterial antigen treated Cal 27. After this treatment, the protein extraction from the treatment conditions and the array procedure was followed according to the manufacturer’s instruction. Details are available in (Supplementary Notes 1 & 2, Supplementary Figure 1, and Supplementary Tables 3 & 4).
V RT-qPCR
SOCS3_F5’GCGCGAAGGCTCCTTTGTG3’;SOCS3_R5’GGGGGGCTGGTCCCGAATC3’;STAT3_F5’GGACATCAGCGGTAAGACCC3’;STAT3_R5’CTCTGGCCGACAATACTTTC3’;EGFR_F5’AGCTACGGGGTGACTGTTTG3’;EGFR_R5’GAACTTTGGGCGACTATCTG3’;GAPDH_F5’GACAGTCAGCCGCATCTTCT3’;GAPDH_R5’ACCAAATCCGTTGACTCCGA3’;PIK3CA_F5’TGGGGATGATTTACGGCAAG3’;PIK3CA_R5’TCCCACACAGTCACCGATTGA3’ were used as primers for the determination of the proliferation-related protein via RT-qPCR (Details are available in Supplementary Note 3 and Supplementary Tables 5-7). Cal 27 cells were plated at a density of 50,000 cells per well in 96-well plates. On subsequent days, bacterial antigen treatments were done. At the completion of the treatment period, total RNA was isolated from the samples with Pure Link RNA mini kit (Invitrogen, Cat #12183018A) and Trizol reagent (Invitrogen, Cat #15596026). The RNA was reverse transcribed by SSIV VILO Master Mix W/EzDNase (Invitrogen, Cat #11766050). qPCR was done using POWRUP SYBR Master Mix (Invitrogen, Cat #A25742). A ViiA 7 qPCR system (Thermo Fisher Scientific) was used to do the comparative Ct (∆∆Ct) analysis.
VI Western Blot
Ten µg of total protein from each bacterial antigen treatment condition was used for Western blotting. We used primary antibodies from R&D Systems: [0.1 µg/ml human/mouse SOCS3 antibody (Cat #MAB5696); 1 µg/ml human PI 3-kinase p110β antibody (Cat #MAB2686); 0.1 µg/ml STAT3 Mouse anti-human, mouse, rat (Cat #MAB1799); 1 µg/ml human EGFR antibody (Cat #AF231), 1:1000 human GAPDH (Cat #2275-PC-020)]. Blots were probed with the respective secondary antibodies from R&D Systems [1:1500 anti-mouse IgG HRP conjugate (Cat #HAF018); 1:1500 anti-goat IgG HRP conjugate (Cat #HAF009); 1:1500 anti-rabbit IgG HRP conjugate (Cat #HAF008)]. Details are available in (Supplementary Note 4).
VII Image Analysis
High-resolution images of clonogenic assay, phosphokinase array, and Western blot were acquired using the Bio-Rad Chemidoc MP Imaging system and processed using Image Lab version 6.0.0 build 25, © 2017 (Bio-Rad Laboratories). For clonogenic assays, the plates were also imaged using a Nikon Coolpix 4500 camera. Colony area intensity, the total number of colonies and densitometric analysis of the images were carried out with plugins for the colony area, particle analysis, and microarray of ImageJ 1.52a (National Institutes of Health; Java 1.8.0_112; 64 bit) (Link). The band density of the loading control protein (untreated Cal 27) and GAPDH (internal control) were used to normalise the protein expression in the treatments.
VIII Statistical Analysis
All statistical analyses were performed using GraphPad Prism® version 5.02. The data were tested for normality of distribution using the D'Agostino & Pearson omnibus normality test. One-way ANOVA with Tukey's Multiple Comparison Test was performed to determine the relative gene and protein expressions related to tumor proliferation and suppression, and metabolic activity/viability/colony formation responses of Cal 27, following bacterial antigen stimulation. The probability threshold of p ≤ 0.05 was considered statistically significant.
Results
I Cal 27 Showed High Bacterial Stimulation Index
Preliminary stimulation studies of LPS and LTA had a negligible effect on SCC4 and SCC9 as determined by MT Glo assay (data not shown). After 72 hr of bacterial antigen stimulation, Cal 27 showed high LPS and LTA stimulation, compared to SCC25 and the normal oral cell line (p ≤ 0.001) (Figure 1). SCC25 was slightly stimulated by LPS but not by LTA. Based on these results, SCC25 was not considered for further investigations and the rest of the assays were solely on Cal 27. Preliminary studies of LPS and LTA had negligible effects on SCC4 and SCC9 (data not shown). Based on these results, SCC25 was not considered for further investigations and the rest of the assays were solely on Cal 27.
Figure 1: Bacterial antigen stimulation effects of oral cancer cells compared to normal oral cells. Results are shown for a) LPS and b) LTA. Biological replicates were obtained from three cell passages and totalled nine. ***indicates statistically significant p ≤ 0.001 for n = 9, where n represents biological replicates. Error bars represent the standard error of the mean.
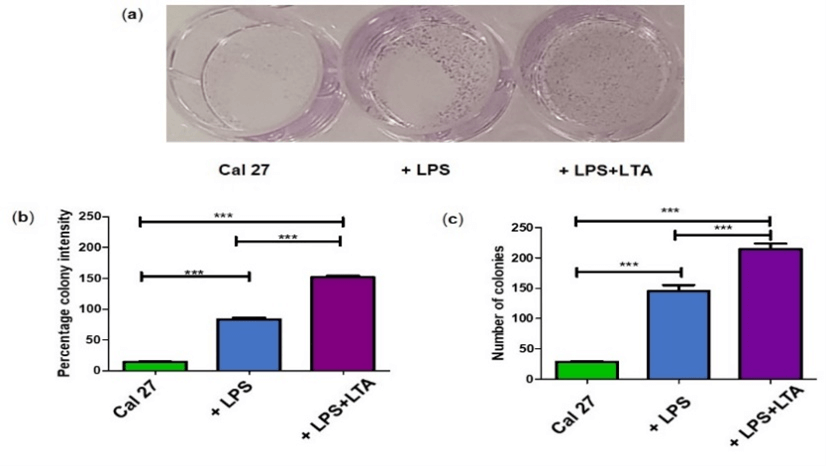
Figure 2: Clonogenic assay of drug combinations after 72 hours stimulation. a) Image of clonogenic assay under different drug conditions; b) Percentage colony intensity of Cal 27 v. different bacterial antigens; c) Number of colonies of Cal 27 v. different bacterial antigens. Biological replicates were obtained from three cell passages and totalled nine.
***indicates statistically significant p ≤ 0.001 for n = 9, where n represents biological replicates. Error bars represent the standard error of the mean.
Figure 3: Comparison of the relative mean pixel density of different kinases under different conditions. Cal 27 after 72 hours LPS stimulation v. Cal 27 after 72 hours LTA stimulation v. Cal 27 after 72 hours LPS+LTA stimulation; at n = 2; Here 'n' represents biological replicates. Biological replicates were obtained from one cell passages and totalled two.
p53: Tumor Suppressor Protein; Chk-2: Checkpoint Kinase 2; p38 α: Mitogen-Activated Protein Kinase 14; GSK3α/β: Glycogen Synthase Kinase 3α/β; GSK3β: Glycogen Synthase Kinase 3β; HSP 60: Heat Shock Protein 60; Yes: Non-Receptor Tyrosine Kinases; Src: Proto-Oncogene Tyrosine-Protein Kinase; EGFR: Epidermal Growth Factor Receptor; eNOS: Endothelial Nitric Oxide Synthase; ERK 1/2: Extracellular Receptor Kinase; STAT3: Signal Transducer and Activator of Transcription 3; RSK 1/2: p90 Ribosomal S6 Kinases.
II LPS+LTA Resulted in Higher Colony Intensity and Number Compared to LPS Stimulation Alone
The percentage colony intensity and the number of colonies estimated by image analysis (Figure 2a) were significantly higher when Cal 27 cells were stimulated with LPS (p ≤ 0.001) (Figures 2b & 2c). Stimulating cells with combined LPS+LTA resulted in even greater colony intensity and more colonies (p ≤ 0.001) (Figures 2b & 2c), confirming our previous finding that bacterial antigens stimulate Cal 27 cells, and this stimulation is greater when both Gram-positive and Gram-negative antigens are combined.
III Combined LPS+LTA Affect Phosphorylated Tumor Suppressor and Proliferation-Related Proteins from Phosphokinase Array Experiment
Phosphokinase array results were analysed by densitometry, to provide relative phosphoprotein expression under treatment conditions of LPS, LTA and LPS+LTA, compared to untreated Cal 27 cells (Relative mean pixel density 1.0), shown in (Figure 3). Tumor suppressor proteins, p53, Chk-2and p38α, were inactivated by LPS+LTA treatment, while HSP 60 remained unaffected compared to LPS. Tumor enhancers, EGFR, β-catenin, Src, ERK 1/2, eNOS, GSKα/β, GSK3β and STAT3, showed diminished phosphorylation in Cal 27 after combined LPS+LTA antigen stimulation while Yes and RSK 1/2 were unaffected compared to LTA. LTA stimulation showed an increase in the phosphorylation of eNOS, GSK3β, GSK3α/β, STAT3, Src, β-catenin, HSP 60 and Chk2 phosphorylation, relative to LPS stimulated cells.
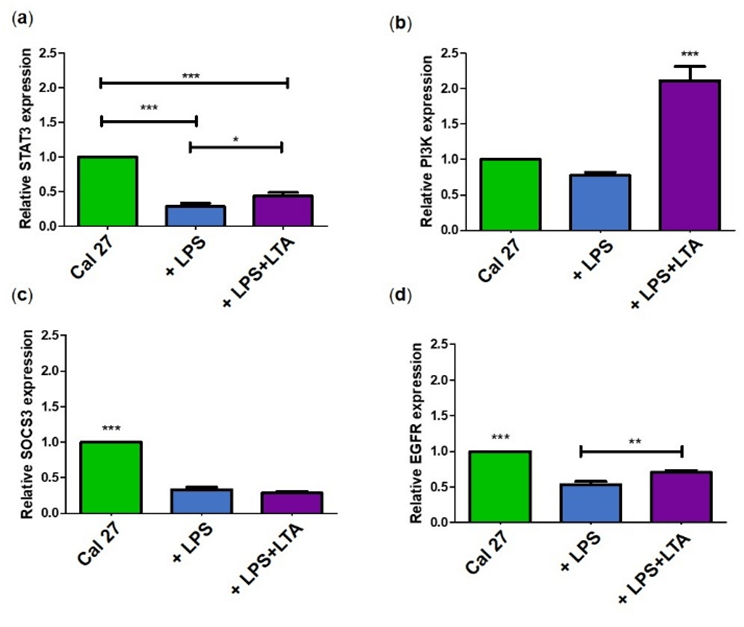
Figure 4: Relative gene expression of different tumor suppressor and proliferation-related proteins in Cal 27. a) Relative STAT3 expression under different conditions; b) Relative PI3KCA expression under different conditions; c) Relative SOCS3 expression under different conditions; d) Relative EGFR expression under different conditions.
***p ≤ 0.001, *p ≤ 0.05 are statistically significant at n = 9. Error bars represent the standard error of the mean. Here 'n' represents biological replicates. All the protein expressions are relative to Cal 27. Biological replicates were obtained from three cell passages and totalled nine.
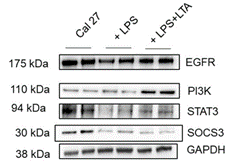
Figure 5: Western blot images of Cal 27 under different treatment conditions. 10-well Western blot image showing bands of different tumor suppressor and proliferation-related proteins in Cal 27 (untreated/control), after LPS stimulation, and after LPS+LTA stimulation.
IV Combined LPS+LTA Resulted in Lowering of Tumor Suppressor and Higher Proliferation-Related Protein
Bacterial antigen treated cells also showed significant lower STAT3 relative gene expression (p ≤ 0.05) (Figure 4a). Similarly, bacterial antigen stimulation resulted in significantly lower EGFR and PIK3CA relative gene expression than unstimulated Cal 27 (p ≤ 0.001) (Figures 4b & 4d) whereas relative gene expression of SOCS3 was highest when Cal 27 were stimulated with LPS (p ≤ 0.001) (Figure 4c). Analysis of the Western blot images (Figure 5) showed that STAT3, SOCS3 and EGFR protein expressions were significantly higher in Cal 27 without bacterial antigen stimulation than bacterial antigen-stimulated cells (p ≤ 0.001) (Figures 6a, 6c & 6d). Also, STAT3 protein expression was significantly higher in Cal 27 after LPS+LTA than after stimulation with LPS (p ≤ 0.05) (Figure 6a). PI3K protein expression was significantly higher after stimulation with LPS+LTA than with no bacterial antigen stimulation or after stimulation with LPS alone (p ≤ 0.001) (Figure 6b). Also, EGFR protein expression was significantly higher in Cal 27 after stimulation of LPS+LTA than after stimulation of LPS (p ≤ 0.005) (Figure 6d).
Figure 6: Western blot analysis of different tumor suppressor and proliferation-related proteins in Cal 27. a) Relative STAT3 expression under different conditions; b) Relative PI3K expression under different conditions; c) Relative SOCS3 expression under different conditions; d) Relative EGFR expression under different conditions.
***p ≤ 0.001, **p ≤ 0.005, *p ≤ 0.05 are statistically significant at n = 6. Error bars represent the standard error of the mean. Here 'n' represents biological replicates. All the protein expressions are relative to Cal 27. Biological replicates were obtained from two cell passages and totalled six.
Discussion
Both LPS and LTA individually stimulate Cal 27 cells but not SCC25, with negligible effect on SCC4 and SCC9. Therefore, this study focussed on Cal 27. Based on the results of Cal 27 MT Glo and clonogenic assays, the LPS+LTA combination has a significant effect than LPS stimulation, supporting bacterial biofilm as plaque may result in increased progression of oral cancer than anaerobic bacteria alone. This result was obtained in the complete absence of immune cells, unlike the Utispan et al. study [3].
While exploring the reason behind the synergistic effect of the combined bacterial antigens, LPS+LTA stimulation resulted in lower levels of phosphorylation of p53 (S15, S46, S392), Chk2 (T68), β-catenin, p38α (T180, Y182), and GSK 3α/β (S21, S9) than it did after stimulation with either LPS or LTA alone. This likely indicates downregulation of tumor suppressor protein activity of p53, p38α and Chk2 (major activator of p53), which are activated by phosphorylation; and downregulation of inactivated β-catenin, which is inactivated by phosphorylation [8-11]. Also, lower levels of phosphorylated (activated) GSK 3α/β and GSK 3β (compared to LTA stimulation) likely reflects the activity of GSK required for binding to inactivated β-catenin for β-catenin degradation and ubiquitination [11]. Also, downregulation of phosphorylated HSP 60 (compared to LTA stimulation), which has previously been shown to have a tumor suppressor-like function by inducing cell differentiation and inhibiting invasion in hepatocellular carcinoma, may further explain the current findings for Cal 27 [12]. However, phosphorylation of other proliferation-related proteins such as Yes-associated proteins (Y426), Src (Y419), EGFR (Y1086), eNOS (S1177), ERK 1/2 (T202/Y204 and T185/Y187) and STAT3 (Y705) was higher in untreated cells than in LPS+LTA stimulated cells. This finding is paradoxical and may require further investigation to explain the reduction in the abovementioned proliferation-related protein, although it has previously been shown that phosphorylation at Y705 of STAT3 may inactivate STAT3 by co-deactivation of S727 of STAT3 [13].
Contrastingly, LTA induced proliferation (that was found in the project) may be due to the phosphorylation of HSP60 (Co-expression of HSP60 and nuclear β catenin), Src and GSK3β. RSK 1/2 has previously been seen to promote invasion and metastasis of head and neck squamous cell carcinoma cells [14]. RSK 1/2 in its phosphorylated (activated) form was seen to be at higher levels in Cal 27 following LPS stimulation than in either after LTA or LPS+LTA stimulation. This may suggest that proliferation caused by LPS stimulation of Cal 27 using RSK 1/2 uses pathways other than Ras/MAPK/ERK 1/2 or that it occurs via upregulation resulting from a positive feedback loop involving other unknown pathways. Paradoxically, EGFR, eNOS, ERK 1/2, Src and RSK 1/2 expression associated with proliferation was found to be lowest after LPS+LTA stimulation, which showed the highest proliferation.
The relative gene expression of SOCS3 and PI3KCA were higher in Cal 27 whether untreated or following LPS stimulation than other treatment conditions. However, Western blot analysis, showed that SOCS3 protein expression was lowest and PI3K expression highest after LPS+LTA stimulation compared with untreated and LPS stimulated cells. This may indicate the lack of translation of SOCS3 and PI3K proteins [15]. Since SOCS3 is known to be a tumor suppressor protein and PI3K is a tumor proliferation-related protein, this result further supports the finding that Cal 27 after LPS+LTA stimulation produced the highest proliferation among all treatment conditions [16, 17]. Similar to the proteome profiling results, Western blot showed lower STAT3 and EGFR expression in Cal 27 after LPS+LTA stimulation compared with untreated cells. This may be because of the lower expression of SOCS3, which has been reported to have a positive feedback loop with STAT3 for Cal 27 [18].
The Western blot revealed lower EGFR protein expression in Cal 27 after LPS stimulation than in untreated cells. In contrast, there was no difference among the two groups in expression levels of phosphorylated EGFR. This implies a difference in phosphorylated (activated) and total protein expression [19]. Therefore, total protein expression does not predict the activity of proliferation proteins in response to the bacterial antigen. Also, the tyrosine 1086 phosphorylation site of EGFR was examined in this study, but other EGFR phosphorylation sites might be activated more in Cal 27 after LPS stimulation [20].
PI3K expression was higher in the LPS+LTA stimulated cells than in untreated and LPS stimulated cells. These findings are in line with previous findings that the PI3K signalling pathway is activated by LTA [21]. Interestingly, the phosphokinase array did not show any appreciable expression of Akt 1/2/3 (serine 473, threonine 308) under any treatment condition, although expression of PI3K was detected in both the RT-qPCR and Western blot. This likely indicates that PI3K induced proliferation of Cal 27, rather than activating the downstream Akt pathway [22]. The project used LTA and LPS to simulate Gram-positive and Gram-negative bacterial infection, respectively. However, components of bacteria other than their bacterial cell wall antigens may be responsible for the progression of oral squamous cell carcinomas. Future studies on oral cancer cell lines, other than SCC4, SCC9, SCC25 and Cal 27 used in this study can give a broader idea regarding the effect of bacterial antigen on oral cancer cells.
Acknowledgement
The project was funded by Sydney Vital Translational Cancer Research Award Round 9 and International Macquarie University Research Excellence Scholarship (iMQRES). The oral cancer cells were kindly donated by Dr. Charbel Darido, Peter MacCallum Cancer Research Centre, University of Melbourne, Australia.
Ethical Approval
This study was approved by the Macquarie University Biosafety Committee (“Mammalian Cell Culture” 5215).
Conflicts of Interest
None.
Article Info
Article Type
Research ArticlePublication history
Received: Mon 11, Jan 2021Accepted: Tue 26, Jan 2021
Published: Mon 08, Feb 2021
Copyright
© 2023 Shoba Ranganathan. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Hosting by Science Repository.DOI: 10.31487/j.DOBCR.2021.01.03
Author Info
Rajdeep Chakraborty Honghua Hu Abubakar Siddiq Mangani Karen Vickery Shoba Ranganathan
Corresponding Author
Shoba RanganathanDepartment of Molecular Sciences, Macquarie University, Sydney, New South Wales, Australia
Figures & Tables
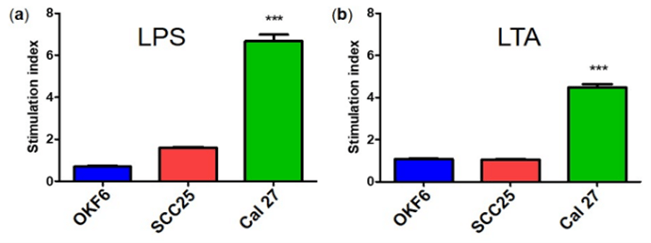
***indicates statistically significant p ≤ 0.001 for n = 9, where n represents biological replicates. Error bars represent the standard error of the mean.

p53: Tumor Suppressor Protein; Chk-2: Checkpoint Kinase 2; p38 α: Mitogen-Activated Protein Kinase 14; GSK3α/β: Glycogen Synthase Kinase 3α/β; GSK3β: Glycogen Synthase Kinase 3β; HSP 60: Heat Shock Protein 60; Yes: Non-Receptor Tyrosine Kinases; Src: Proto-Oncogene Tyrosine-Protein Kinase; EGFR: Epidermal Growth Factor Receptor; eNOS: Endothelial Nitric Oxide Synthase; ERK 1/2: Extracellular Receptor Kinase; STAT3: Signal Transducer and Activator of Transcription 3; RSK 1/2: p90 Ribosomal S6 Kinases.

***p ≤ 0.001, *p ≤ 0.05 are statistically significant at n = 9. Error bars represent the standard error of the mean. Here 'n' represents biological replicates. All the protein expressions are relative to Cal 27. Biological replicates were obtained from three cell passages and totalled nine.
***p ≤ 0.001, **p ≤ 0.005, *p ≤ 0.05 are statistically significant at n = 6. Error bars represent the standard error of the mean. Here 'n' represents biological replicates. All the protein expressions are relative to Cal 27. Biological replicates were obtained from two cell passages and totalled six.
References
- Coelho KR (2012) Challenges of the oral cancer burden in India. J Cancer Epidemiol 2012: 701932. [Crossref]
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA et al. (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68: 394-424. [Crossref]
- Utispan K, Pugdee K, Koontongkaew S (2018) Porphyromonas gingivalis lipopolysaccharide-induced macrophages modulate proliferation and invasion of head and neck cancer cell lines. Biomed Pharmacother 101: 988-995. [Crossref]
- Karpinski TM (2019) Role of Oral Microbiota in Cancer Development. Microorganisms 7: 20. [Crossref]
- Matsumoto K, Umitsu M, De Silva DM, Roy A, Bottaro DP (2017) Hepatocyte growth factor/MET in cancer progression and biomarker discovery. Cancer Sci 108: 296-307. [Crossref]
- Sugiyama A, Arakaki R, Ohnishi T, Arakaki N, Daikuhara Y et al. (1996) Lipoteichoic acid and interleukin 1 stimulate synergistically production of hepatocyte growth factor (scatter factor) in human gingival fibroblasts in culture. Infect Immun 64: 1426-1431. [Crossref]
- Li H, Wawrose JS, Gooding WE, Garraway LA, Yan Lui VW et al. (2014) Genomic analysis of head and neck squamous cell carcinoma cell lines and human tumors: a rational approach to preclinical model selection. Mol Cancer Res 12: 571-582. [Crossref]
- DeNicola GF, Martin ED, Chaikuad A, Bassi R, Clark J et al. (2013) Mechanism and consequence of the autoactivation of p38alpha mitogen-activated protein kinase promoted by TAB1. Nat Struct Mol Biol 20: 1182-1190. [Crossref]
- Jenkins LM, Durell SR, Mazur SJ, Appella E (2012) p53 N-terminal phosphorylation: a defining layer of complex regulation. Carcinogenesis 33: 1441-1449. [Crossref]
- Xu X, Tsvetkov LM, Stern DF (2002) Chk2 activation and phosphorylation-dependent oligomerization. Mol Cell Biol 22: 4419-4432. [Crossref]
- Liu C, Li Y, Semenov M, Han C, Baeg GH et al. (2002) Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 108: 837-847. [Crossref]
- Zhang J, Zhou X, Chang H, Huang X, Guo X et al. (2016) Hsp60 exerts a tumor suppressor function by inducing cell differentiation and inhibiting invasion in hepatocellular carcinoma. Oncotarget 7: 68976-68989. [Crossref]
- Yang J, Kunimoto H, Katayama B, Zhao H, Shiromizu T et al. (2020) Phospho-Ser727 triggers a multistep inactivation of STAT3 by rapid dissociation of pY705-SH2 through C-terminal tail modulation. Int Immunol 32: 73-88. [Crossref]
- Kang S, Elf S, Lythgoe K, Hitosugi T, Taunton J et al. (2010) p90 ribosomal S6 kinase 2 promotes invasion and metastasis of human head and neck squamous cell carcinoma cells. J Clin Invest 120: 1165-1177. [Crossref]
- Gsponer J, Babu MM (2012) Cellular strategies for regulating functional and nonfunctional protein aggregation. Cell Rep 2: 1425-1437. [Crossref]
- Inagaki Ohara K, Kondo T, Ito M, Yoshimura A (2013) SOCS, inflammation, and cancer. JAKSTAT 2: e24053. [Crossref]
- Arcaro A, Guerreiro AS (2007) The phosphoinositide 3-kinase pathway in human cancer: genetic alterations and therapeutic implications. Curr Genomics 8: 271-306. [Crossref]
- Carow B, Rottenberg ME (2014) SOCS3, a Major Regulator of Infection and Inflammation. Front Immunol 5: 58. [Crossref]
- Hunter T (2000) Signaling--2000 and beyond. Cell 100: 113-127. [Crossref]
- Tong J, Taylor P, Peterman SM, Prakash A, Moran MF (2009) Epidermal growth factor receptor phosphorylation sites Ser991 and Tyr998 are implicated in the regulation of receptor endocytosis and phosphorylations at Ser1039 and Thr1041. Mol Cell Proteomics 8: 2131-2144. [Crossref]
- Kao SJ, Lei HC, Kuo CT, Chang MS, Chen BC et al. (2005) Lipoteichoic acid induces nuclear factor-kappaB activation and nitric oxide synthase expression via phosphatidylinositol 3-kinase, Akt, and p38 MAPK in RAW 264.7 macrophages. Immunology 115: 366-374. [Crossref]
- Faes S, Dormond O (2015) PI3K and AKT: Unfaithful Partners in Cancer. Int J Mol Sci 16: 21138-21152. [Crossref]
